Research projects
We are working on a range of problems that include foundational methods and automatic control techniques for synthetic biology, as well as study of the dynamics of complex biological networks.
Foundational methods for genetic circuit design and control
Host-aware synthetic biology
 When bacteria are engineered to express a foreign protein, this adds an extra 'load' or 'burden' on these cells by taking away resources that would naturally be used for growth. Bacteria have natural internal mechanisms to sense that they are burdened and adapt to deal with the new load. Discovering these mechanisms is not only important at a fundamental level but also to develop a new generation of "host-aware" synthetic biology systems that can sense the load they impose on their cell and, in response, automatically adjust their level of genetic expression to regulate this load and ensure a good balance between synthetic function and tolerable host cell burden. At their core, "host-aware" synthetic biology systems rely on the design, fine-tuning, and implementation in living cells of general biomolecular feedback controllers. The use of feedback system embedded in living cells allows to build synthetic biology systems that robustly maintain performance in the presence of perturbations and uncertainties, which ultimately improves the reliability and predictability of synthetic biology designs.
When bacteria are engineered to express a foreign protein, this adds an extra 'load' or 'burden' on these cells by taking away resources that would naturally be used for growth. Bacteria have natural internal mechanisms to sense that they are burdened and adapt to deal with the new load. Discovering these mechanisms is not only important at a fundamental level but also to develop a new generation of "host-aware" synthetic biology systems that can sense the load they impose on their cell and, in response, automatically adjust their level of genetic expression to regulate this load and ensure a good balance between synthetic function and tolerable host cell burden. At their core, "host-aware" synthetic biology systems rely on the design, fine-tuning, and implementation in living cells of general biomolecular feedback controllers. The use of feedback system embedded in living cells allows to build synthetic biology systems that robustly maintain performance in the presence of perturbations and uncertainties, which ultimately improves the reliability and predictability of synthetic biology designs.
Using a burden-based biomolecular controller, we engineered the first example of 'host-aware' synthetic biology system and showed that such systems are modular, tuneable and portable. Their use opens exciting possibilities for synthetic biology in particular, where the implementation within a single cell of large genetic constructs needs to be reliable and genetically stable, and done in a way that does not affect the cell's performance. The method we used and the results are presented in our 2018 Nature Methods paper in Collaboration with the Ellis lab.
Quantification of gene expression burden
 The introduction of a synthetic genetic circuit into a host chassis cell is often non-trivial in terms of predicting the behaviour of both. Expression from a synthetic circuit uses up the same cellular resources and machinery required for the growth and maintenance of the host cell. By sequestering these resources away from native cellular functions, a synthetic system can have detrimental effects on the cell, which in turn feedback upon the synthetic system behaviour. As the requirement for larger and more complex synthetic circuits increases, it is important to understand the mechanisms through which host cell and circuit interact. Our research combines wet-lab and modelling work to understand, monitor and quantify the burden a synthetic circuit places upon a cell with the aim of developing a predictive system for the behaviour of both.
The introduction of a synthetic genetic circuit into a host chassis cell is often non-trivial in terms of predicting the behaviour of both. Expression from a synthetic circuit uses up the same cellular resources and machinery required for the growth and maintenance of the host cell. By sequestering these resources away from native cellular functions, a synthetic system can have detrimental effects on the cell, which in turn feedback upon the synthetic system behaviour. As the requirement for larger and more complex synthetic circuits increases, it is important to understand the mechanisms through which host cell and circuit interact. Our research combines wet-lab and modelling work to understand, monitor and quantify the burden a synthetic circuit places upon a cell with the aim of developing a predictive system for the behaviour of both.
As part of this research, we have developed a simulation platform based on a model of translation under finite ribosome sharing between the cell and the synthetic biology circuits it hosts. This platform was used to generate the results presented in our 2015 Nature Methods paper. You can download here, the corresponding python simulation and matlab visualisation code to run your own simulations.
Easily Tunable Synthetic Biology
 The ability to easily tune the dynamic properties of engineered biomolecular and genetic systems facilitates their use in different contexts and for different purposes. Genetic oscillators were some of the first bio-circuits constructed in synthetic biology and have already been used in various applications. Current designs do not allow to control and tune the amplitude without affecting the frequency of the periodic oscillation, or vice versa. Using computer modelling we investigate the mechanisms through which amplitude and frequency are coupled in current genetic oscillators, and, based on this, propose new engineering design principles to control independently amplitude and frequency and over extended ranges in response to external tuning inputs.
The ability to easily tune the dynamic properties of engineered biomolecular and genetic systems facilitates their use in different contexts and for different purposes. Genetic oscillators were some of the first bio-circuits constructed in synthetic biology and have already been used in various applications. Current designs do not allow to control and tune the amplitude without affecting the frequency of the periodic oscillation, or vice versa. Using computer modelling we investigate the mechanisms through which amplitude and frequency are coupled in current genetic oscillators, and, based on this, propose new engineering design principles to control independently amplitude and frequency and over extended ranges in response to external tuning inputs.
Oscillators are fundamental components in electronic devices like radios and computers. Taking inspiration from tuneable periodic signal generators widely found in electronics, we are interested in creating the equivalent in biology. Such genetic oscillators are useful for controlling the behaviour of biological systems with high accuracy and also have potential applications in biosensing, metabolic engineering, and periodic production and delivery of therapeutic molecules. We have have employed computational modelling to analyse genetic oscillators that were proposed previously and based on this we have proposed new designs for the construction of genetic oscillators that can serve as periodic signal generators in biology. The results of our computational modelling study on new architectures for orthogonally tunable oscillators are presented in our 2018 Cell Systems paper, with a short layman introduction by Prof Mary Dunlop.
Using a population of engineered cells to regulate extracellular chemical concentration
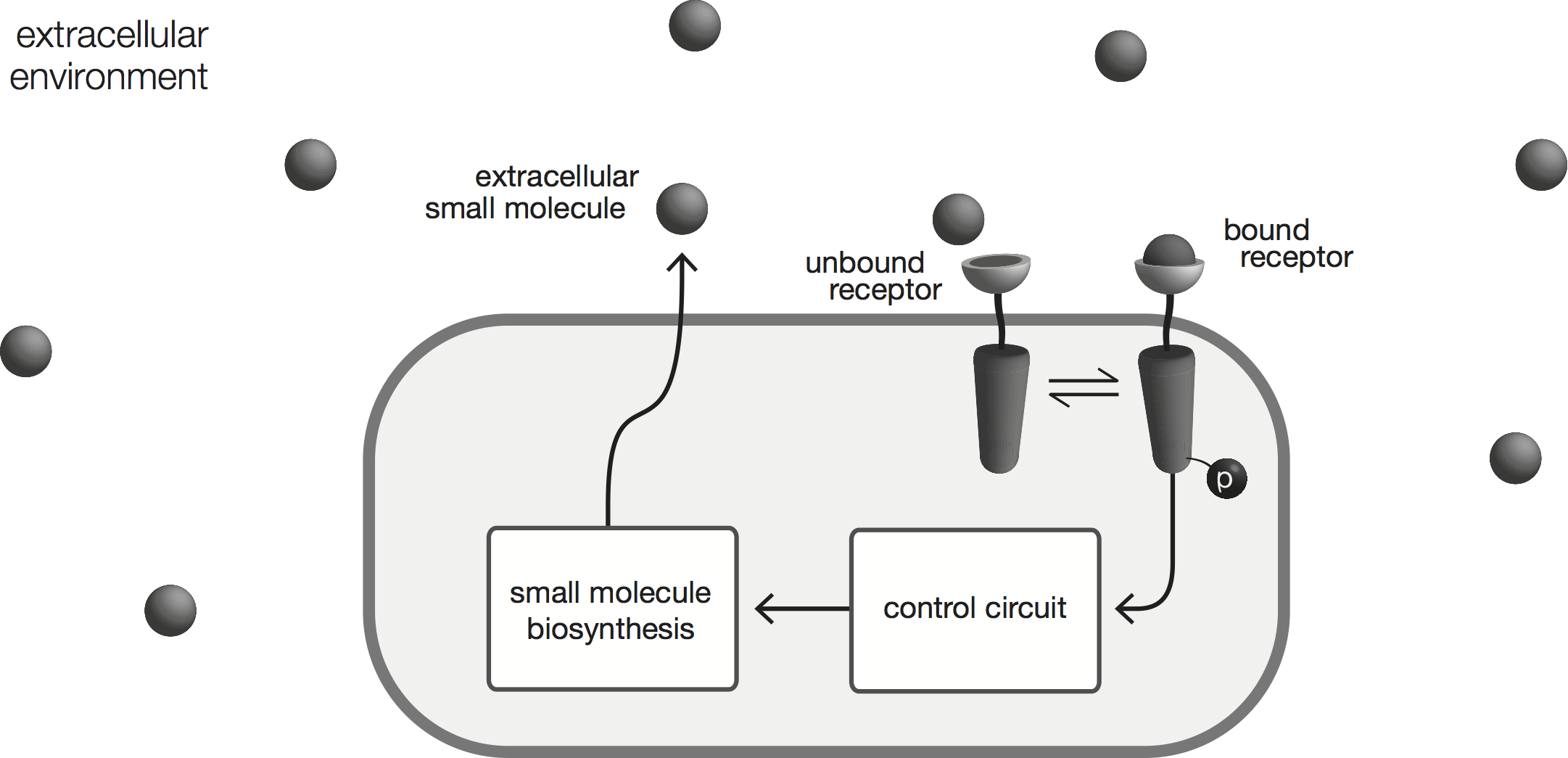 This research focuses on engineering cells to actively sense and regulate the extracellular concentration of a small molecule, using synthetic gene circuits designed to implement control engineering strategies fundamental to robust regulation.
This research focuses on engineering cells to actively sense and regulate the extracellular concentration of a small molecule, using synthetic gene circuits designed to implement control engineering strategies fundamental to robust regulation.
As part of this research, we have organised an International Workshop "On the Prospects for Controllable Cell-Based Therapies" at City University London, 22-23 February 2016.
Engineering biological robustness using ribozyme feedback control
Synthetic biology seeks to engineer biological systems through design of biological pathways and circuits and seeks to produce useful chemical products in a sustainable manner. One problem with producing highly engineered systems is that the system itself may become more fragile the more it is modified. This means that typically biologically engineered systems will not robustly maintain their output in response to external factors or mutations.
To address this issue we aim to implement dynamic control within synthetic circuits. Dynamic control allows systems to respond to external perturbations that may otherwise disrupt normal function. To implement dynamic control synthetically we have chosen to engineer a ribozyme to recognize a small metabolite molecule. This will allow us to generate a feedback circuit whereby the amount of enzyme in the cell is directly regulated by the small metabolite.
Transcriptional control circuits for metabolic networks
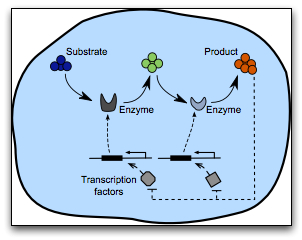 This research investigates the potential of using synthetic transcriptional circuits for the control of bacterial metabolism. The engineered synthetic transcriptional circuits modulate metabolic activity so as to match cellular demands and act by adjusting gene transcription via transcription factors that are responsive to changes in the metabolic product of interest. These controllers are intracellular, and therefore can operate autonomously without human intervention. We are benchmarking different controller "wirings" and assessing their quality with respect to well-defined criteria such as stability, response time, and oscillatory behaviour. This research aims at providing a valuable model-based understanding of which controllers hold promise for future wet-lab implementations, and which ones should be discarded from the outset.
This research investigates the potential of using synthetic transcriptional circuits for the control of bacterial metabolism. The engineered synthetic transcriptional circuits modulate metabolic activity so as to match cellular demands and act by adjusting gene transcription via transcription factors that are responsive to changes in the metabolic product of interest. These controllers are intracellular, and therefore can operate autonomously without human intervention. We are benchmarking different controller "wirings" and assessing their quality with respect to well-defined criteria such as stability, response time, and oscillatory behaviour. This research aims at providing a valuable model-based understanding of which controllers hold promise for future wet-lab implementations, and which ones should be discarded from the outset.
External computer-based control of engineered cells and synthetic gene circuits
Data-based optimal control of synthetic biology gene networks
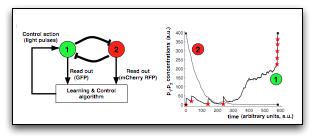 In this research we seek to develop methods and associated algorithms for the optimal control of gene networks without the need for a priori identifying dynamical models for these networks. An outstanding feature of our approach is that the control techniques we are developing are solely based on input-output data readouts from the network, with the goal to provide optimal and robust operation of potentially poorly characterised or unknown networks. We call such an approach data-based optimal control as opposed to more classical model-based approaches. Data-based methods aim to infer/learn a feedback optimal control law from input-output data collected from the system, i.e., from the way the output of the system reacts to various perturbations at the input of the system.
In this research we seek to develop methods and associated algorithms for the optimal control of gene networks without the need for a priori identifying dynamical models for these networks. An outstanding feature of our approach is that the control techniques we are developing are solely based on input-output data readouts from the network, with the goal to provide optimal and robust operation of potentially poorly characterised or unknown networks. We call such an approach data-based optimal control as opposed to more classical model-based approaches. Data-based methods aim to infer/learn a feedback optimal control law from input-output data collected from the system, i.e., from the way the output of the system reacts to various perturbations at the input of the system.
The vision of this research is to provide foundational methods that will enable the "automatic learning" of strategies for optimally controlling gene regulation networks, i.e., the creation of computer based systems that automatically learn input-scheduling strategies so as to optimally control gene regulation networks by interacting with them. The control strategies learned by our methods take into account noise and uncertainties in the data and are developed to be robust with respect to these. Such data-based strategies are analogous to, for example, the way we drive our cars: without any a priori mathematical model of the car behaviour on the road, we can effectively learn how and when to steer, accelerate and break (inputs) based on our observations of the car's position and velocity on the road (outputs) so as to, for example, learn to minimise our lap time around an unknown track using appropriate input scheduling strategies.
The data-based optimal control methods we are currently developing are quite general and thus will be applicable to various optimal control problems arising in the control of biological or technological systems.
You can download PyFQI, our implementation in python of the Fitted Q Iteration Algorithm. The corresponding PyFQI-mini-manual can be downloaded here.
Light-based control of synthetic gene circuits
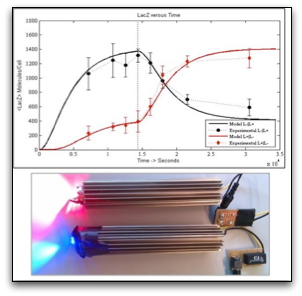 Utilising biological light sensors to gain control over transcriptional regulation was realised through years of research on the structural nature of the light-harvesting protein domains, encountered mainly in photosynthetic organisms. The Cph8 chimera introduced by Levskaya et al. (2005), was one of the first examples on how light can provide a non-invasive, cheap and reversible induction scheme. These advantages, absent in chemical inducers, hold promising potentials for developing a framework for gaining temporal and even spatial control over cellular metabolism. Our work is focused on the experimental characterisation and mathematical modeling of the dynamics of light responsive systems and on optimising them for a robust and predictable transcriptional photo-regulation. Next we aim to couple genetic oscillators with the light input modules in order to gain control over the frequency and amplitude of the oscillation through light.
Utilising biological light sensors to gain control over transcriptional regulation was realised through years of research on the structural nature of the light-harvesting protein domains, encountered mainly in photosynthetic organisms. The Cph8 chimera introduced by Levskaya et al. (2005), was one of the first examples on how light can provide a non-invasive, cheap and reversible induction scheme. These advantages, absent in chemical inducers, hold promising potentials for developing a framework for gaining temporal and even spatial control over cellular metabolism. Our work is focused on the experimental characterisation and mathematical modeling of the dynamics of light responsive systems and on optimising them for a robust and predictable transcriptional photo-regulation. Next we aim to couple genetic oscillators with the light input modules in order to gain control over the frequency and amplitude of the oscillation through light.
Identification and control of nonlinear dynamical systems and complex dynamical networks
Control Engineering of Biological Systems for Reliable Synthetic Biology Applications (COSY-BIO)
We are working as part of the EU H2020 FET-OPEN RIA grant 766840 COSY-BIO, on developing new control engineering methods and tools for reliable synthetic biology applications. This is large collaborative work between 8 leading institutions in Europe: TIGEM (Italy), Imperial (UK), University of Edinburgh (UK), University of Bristol (UK), ETH Zurich (Switzerland), Institut Pasteur (France), University Paris-Diderot (France), Instituto Italiano di Tecnologia (Italy). Please see the COSY-BIO website for more information.
Automatic identification of nonlinear ordinary differential equation models from time-series data
Prediction and control of behaviour and abnormalities in any complex dynamical network, and in particular in those encountered in biology require the development of multivariate predictive models that integrate large dataset from different sources. Although, a large amount of data are being collected on a daily basis, very few methods allow the automatic creation from these data of nonlinear Ordinary Differential Equation models for understanding and (re-)design/control, and an inordinate amount of time is still being spent on the manual aggregation of information and expert development of models that explains these data.
In this context, the problem of reconstruction or identification of biological systems from experimental time series data is of fundamental importance. Yet, the development of general reconstruction techniques remains challenging, especially for nonlinear system identification. We are currently developing new methods to identify both parametric structure and parameter values in nonlinear Ordinary Differential Equation models from heterogeneous datasets. Applications of such nonlinear systems identification methods cover fundamental questions in systems biology, synthetic biology (debugging and design of cellular systems) and modelling of complex dynamical networks.
You can download BSID, our Matlab toolbox for nonlinear systems identification from time-series data.
Synchronisation, consensus and collective behaviours in complex dynamical networks
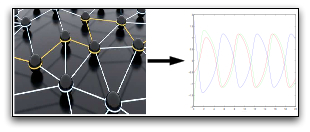 In this research, we investigate questions related to the emergence and control of collective behaviours in complex networks of interconnected dynamical systems. In particular, we focus on questions pertaining to synchronisation and consensus in complex dynamical networks modelling technological systems (e.g., agreement social networks, distributed computing, synchronisation of coupled oscillators) and biological systems (e.g., synchronisation in bacterial populations, circadian rhythms, heart beats, swarming and flocking). We are in particular interested in the application of such decentralised control methods to the control of cell populations so as to force them to cooperatively perform a desired task.
In this research, we investigate questions related to the emergence and control of collective behaviours in complex networks of interconnected dynamical systems. In particular, we focus on questions pertaining to synchronisation and consensus in complex dynamical networks modelling technological systems (e.g., agreement social networks, distributed computing, synchronisation of coupled oscillators) and biological systems (e.g., synchronisation in bacterial populations, circadian rhythms, heart beats, swarming and flocking). We are in particular interested in the application of such decentralised control methods to the control of cell populations so as to force them to cooperatively perform a desired task.